 End-to-end pipeline for mult-scale modelling
End-to-end pipeline for mult-scale modelling
Authors: Nazanin Ebrahimi, Mahyar Osanlouy, Chris Bradley, Fabiana Kubke, Dane Gerneke, Peter Hunter
Publication: iScience. (July 2022)
Dataset: Mendeley Data
Overview
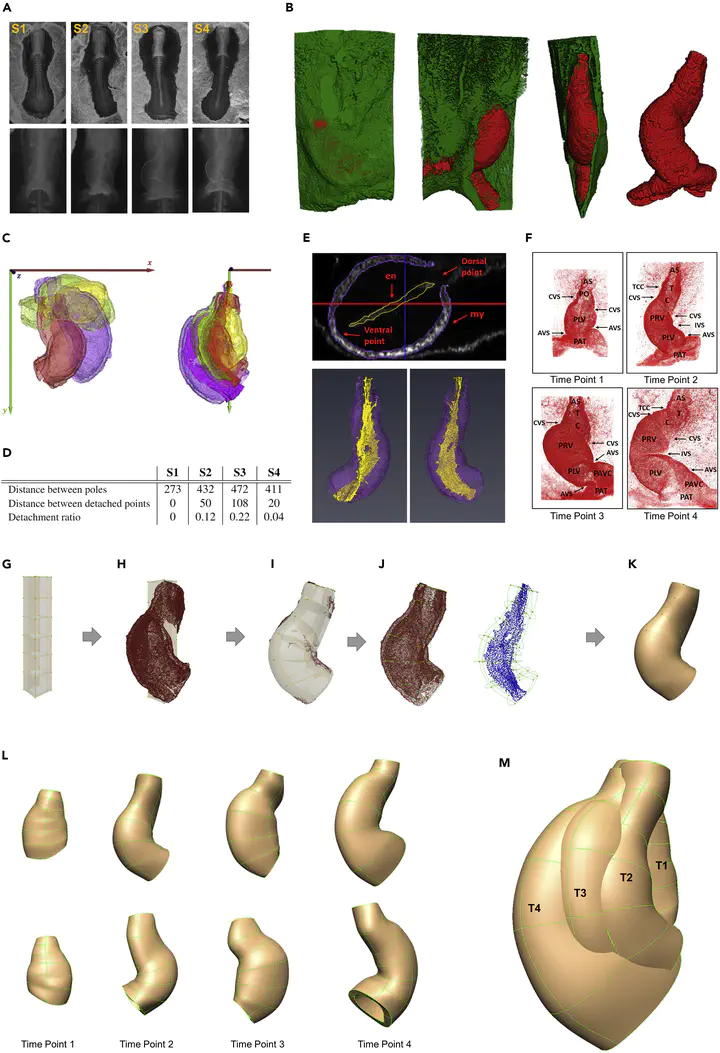
This study presents an innovative hybrid experimental-computational pipeline to investigate the spatiotemporal growth patterns underlying cardiac C-looping in embryonic chick hearts. C-looping is a critical phase in heart development where the straight heart tube transforms into a C-shaped structure, setting the stage for proper chamber formation. Abnormalities during this process are linked to congenital heart defects. The work combines multi-scale imaging, deep learning-based cell segmentation, and biomechanical modeling to bridge cellular dynamics with tissue-level deformations, offering new insights into the mechanisms driving heart morphogenesis.
Key Contributions
- First 3D multi-scale dataset of C-looping hearts, integrating cell-to-organism level data from confocal microscopy and micro-CT.
- Fully automated 3D myocardial cell segmentation using a custom convolutional neural network (CNN), achieving a Dice coefficient of 0.91 ± 0.1.
- Finite Element (FE) biomechanical models capturing spatiotemporal heart geometry changes across four developmental timepoints.
- Variance-driven analysis revealing how inter-cellular space (ICS) and cellular heterogeneity contribute to tissue growth.
- Open-source pipeline for integrating cellular features with tissue-level kinematics, enabling future studies on cardiac morphogenesis.
Methodology
Experimental Workflow
-
Multi-modal imaging:
- Confocal microscopy: Whole-mount staining (WGA, NCAM-1, DAPI) provided 3D cell-resolution images of myocardial cells.
- Micro-CT scanning: Sub-micron resolution imaging contextualized heart geometry within the entire embryo.
- Optical clearing: Enabled high-resolution imaging while preserving 3D architecture.
-
Temporal staging: Embryos were ordered developmentally using anatomical landmarks and dorsal detachment metrics.

Computational Workflow
-
3D Deep Learning for Cell Segmentation:
- A 3D U-Net architecture was trained to segment individual myocardial cells from confocal stacks.
- Achieved 91% Dice similarity compared to manual segmentation, reducing processing time from 8 hours to seconds per 100 cells.
- Addressed memory constraints via a traceable slicing-merging algorithm for large-scale image processing.
-
Finite Element Modeling:
- Anatomically accurate FE meshes were fitted to heart geometries using OpenCMISS.
- RMS fitting error < 3 μm enabled precise representation of tissue deformation.
-
Spatiotemporal Analysis:
- Cellular features (volume, anisotropy, orientation) were mapped onto FE meshes.
- General Linear Model (GLM) linked tissue growth to cellular dynamics (cell number, ICS volume, anisotropy).

Results
-
Differential Growth Patterns:
- Ventral regions showed ~2x higher tissue growth than dorsal regions, driven by increased cell proliferation and ICS expansion.
- Outer curvature cells exhibited circumferential alignment, while inner curvature cells remained isotropic.
-
Role of Inter-Cellular Space (ICS):
- ICS accounted for 30–45% of tissue volume changes, highlighting its role in accommodating growth.
- Ventral ICS volume increased by 62% during bending phases.
-
Variance as a Developmental Signal:
- Ring-shaped variance patterns in cell volume/anisotropy emerged around the outer curvature, suggesting mechanical feedback loops.
- High-variance regions correlated with zones of rapid tissue remodeling.
-
Tissue-Cell Dynamics:
- GLM analysis explained 70% of tissue growth variance, with cell number (β = 0.48, p < 0.001) and ICS (β = 0.32, p < 0.01) as key predictors.
- Cell orientation aligned with tissue deformation vectors during rotation (r > 0.8) but not bending phases.
Implications
- Mechanistic Insights: Challenges the “differential growth hypothesis” by showing ICS and cellular heterogeneity are equally critical to looping.
- Toolkit for Developmental Biology: The pipeline is extensible to other organs/species, enabling studies on how cellular noise shapes tissue patterning.
- Clinical Relevance: Provides a framework to explore how genetic mutations disrupt growth coordination, informing congenital heart defect research.
- AI/ML Impact: Demonstrates the power of deep learning in automating large-scale 3D biological image analysis.
